| Possible Reason | Solution |
| Input amount of nucleic acid is too low |
If
the sample is a co-extracted total nucleic acid, quantification of RNA and DNA
should be performed separately using Qubit, and library amplification should be
carried out according to the recommended cycles in the manual. |
|
During the experiment,
RNase-free consumables must be used, and the operating environment should be
free of RNase contamination, as this can affect the efficiency of RNA reverse
transcription.
|
|
| If the nucleic acid sample contains excessive amounts of chelating agents, guanidine salts, phenol, proteins, ethanol, or other impurities, it can impact the activity of RNA reverse transcriptase and the efficiency of DNA enzymatic digestion. It is recommended refer to Step 7: Post-amplification Clean-up to purify the nucleic acid samples using 2X NadPrep SP Beads, replace the elution reagent with Nuclease Free Water, and then proceed with library preparation. |
The lower limit of mutation detection corresponding to different experimental conditions
|
Library conversion rate |
Sample Input |
|||||||||
|
1 ng |
10 ng |
20 ng |
30 ng |
50 ng |
100 ng |
200 ng |
300 ng |
500 ng |
1000 ng |
|
|
20% |
5.000% |
0.500% |
0.250% |
0.168% |
0.100% |
0.050% |
0.025% |
0.017% |
0.010% |
0.005% |
|
30% |
3.400% |
0.340% |
0.170% |
0.113% |
0.068% |
0.034% |
0.017% |
0.011% |
0.007% |
0.003% |
|
50% |
2.000% |
0.200% |
0.100% |
0.067% |
0.040% |
0.020% |
0.010% |
0.007% |
0.004% |
0.002% |
|
100% |
1.000% |
0.100% |
0.050% |
0.033% |
0.020% |
0.010% |
0.005% |
0.003% |
0.002% |
0.001% |
When the library conversion rate is 50%, 0.005% of the mutations can be detected with a sample input of 400 ng;
②Increase the number of mutation sites
① Increase the sample input:The more sample input , the copies of support low-frequency mutation detection will be more;
② Increase the number of tracked mutations: the higher the number of mutation sites, the probability of detection will be higher.
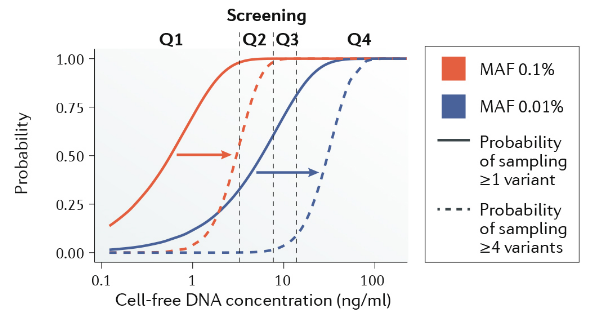 |

|
Van Der Pol Y, Mouliere F. Toward the early detection of cancer by decoding the epigenetic and environmental fingerprints of cell-free DNA[J]. Cancer cell, 2019, 36(4): 350-368.
|
Possible Reason |
Solution |
|
Relatively high starting input |
Quantification by Qubit™ 3.0 is recommended. |
|
Relatively high amplification cycles |
Explore the optimal amplification cycles according to the recommended protocol in the operating guideline. |
|
The elution is not performed according to operational guideline, leading to serious off-target. |
The elution is performed in strict accordance with the requirements of operating guideline. |
|
Possible Reason |
Solution |
|
Inaccurate starting input |
In case of lower actual starting input, precise quantification by qPCR is recommended. |
|
Reaction mixture is not completely transferred out when pipetting. |
Transfer all reaction mixture from each step to a new tube when pipetting. |
|
Some magnetic beads are discarded during hybridization and bead purification |
Avoid too intense operation during the elution, and discard supernatant without losing any magnetic beads. |

