No Fear of Sample Damage—An All-scenario DNA Methylation Solution that Detects 10 Major High-incidence Cancers in A Single Assay!
01 Background
Malignant tumors have become a major global public health challenge; screening and early diagnosis with timely treatment can effectively reduce cancer mortality, and interventions based on “early screening, early detection, early intervention” can significantly increase cure rates and survival rates. Typically, during proliferation or apoptosis, tumor cells release DNA fragments (ctDNA) carrying specific mutations or methylation biomarkers into the bloodstream, forming a “molecular fingerprint” of cancer. Compared with traditional tumor biomarkers, DNA methylation biomarkers offer earlier detection, less invasiveness, and higher precision, making them well suited for multiple scenarios including cancer screening, auxiliary diagnosis, companion diagnostics, monitoring of minimal residual disease (MRD) and recurrence alerts as well as prognostic assessment[1-5]. Studies have shown that combining ctDNA methylation with mutation biomarkers for longitudinal and MRD monitoring can increase MRD detection sensitivity by approximately 25% ~ 36%[6].
However, conventional DNA methylation detection typically relies on sequence conversion, such as bisulfite and enzymatic methods: the former is fast and efficient but prone to DNA degradation and fragmentation, limiting its use to samples with high input; the latter is gentler but more laborious, time-consuming, and less consistently conversion efficient. Therefore, Nanodigmbio has launched an ultra-sensitive solution for synchronous detection of methylation and mutation (μCaler DNA Full Screen System). This solution eliminates the need for sequence conversion, enabling ultra-sensitive detection of both methylation and mutations from limited input samples. It simplifies the workflow while maximizing the retention of original template information and significantly improves coverage depth. Especially in low-abundance signal detection scenarios, the μCaler DNA Full Screen System is poised to become a key tool for early cancer screening and postoperative MRD surveillance.
Further Reading
- No Need for Conversion: Ultra-sensitive Solution for Synchronous Detection of Methylation and Mutation in a Single Reaction Coming Soon!
- How to Achieve Ultra-sensitive Synchronous Detection of Methylation and Mutation without Sequence Conversion?
- How Effective is the Ultra-sensitive Solution for Synchronous Detection of Methylation and Mutation? Let Clinical Samples be the Judge!
To fully exploit this solution’s advantages, Nanodigmbio now offers the μCaler FS EMS+ Panel v1.0, which comprehensively covers key CpG sites across 10 major high-incidence cancers and enables precise detection of methylation states in candidate genes for target cancer types, providing a powerful tool for the clinical application of DNA methylation biomarkers in precision oncology and accelerating the translation of research findings into clinical practice.
02 Solution Overview
2.1 Introduction
μCaler FS EMS+ Panel v1.0 includes selected sites approved by the NMPA and FDA, as well as those reported in literature and patents, totaling 1,783 CpG sites. It covers 10 major high-incidence cancers, including lung cancer, breast cancer, colorectal cancer, prostate cancer, gastric cancer, liver cancer, cervical cancer, bladder cancer, esophageal cancer, and ovarian cancer, involving 163 methylated candidate genes associated with carcinogenesis and tumor suppression. The panel’s target regions cover approximately 0.1 Mb of the human genome, with 9,680 CpG sites, providing strong technical support for comprehensive and accurate methylation early screening and MRD recurrence monitoring.
Table 1. Target cancers and methylation candidate genes covered by the μCaler FS EMS+ Panel v1.0.
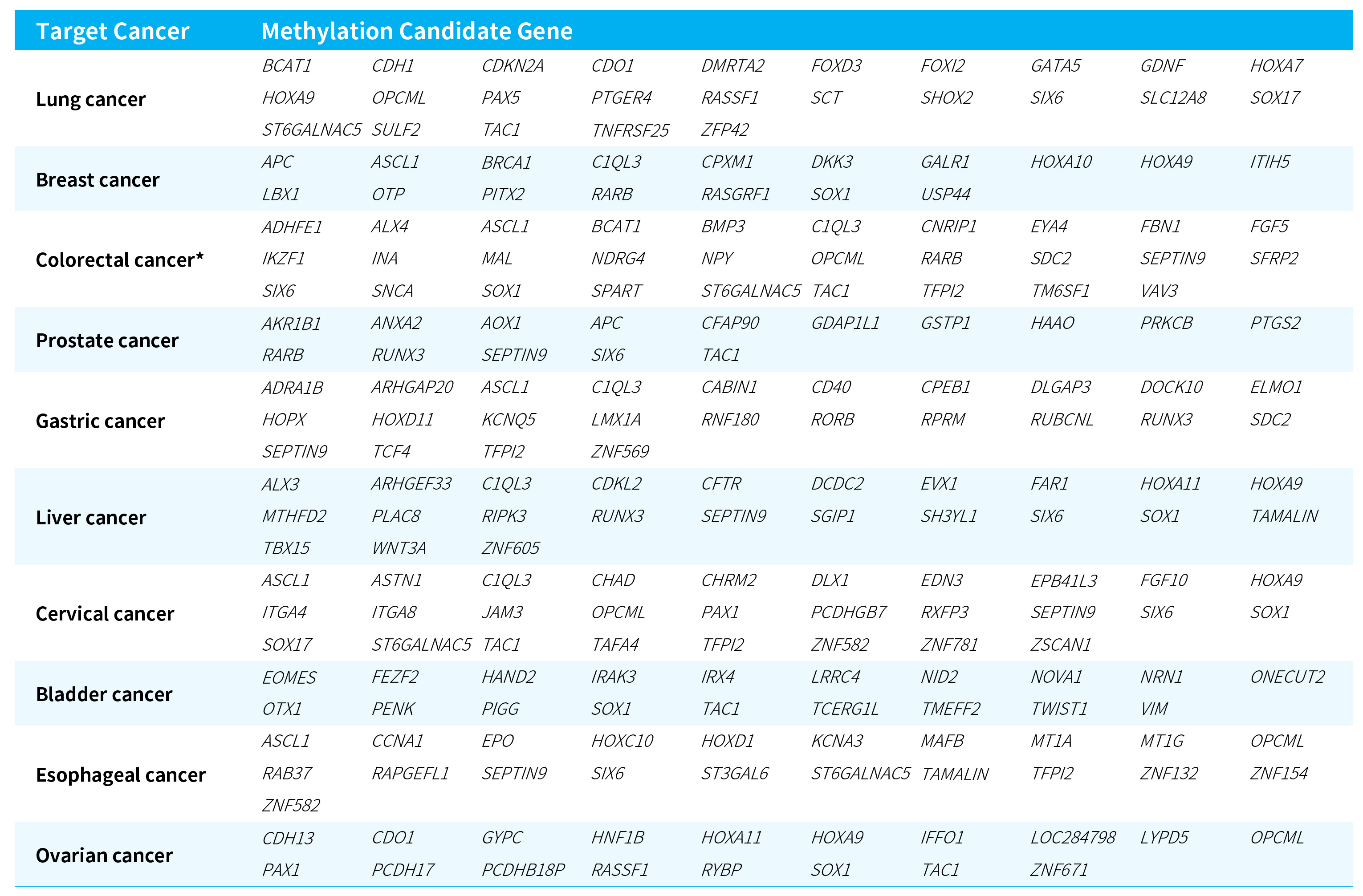
*It also covers intergenic regions not listed in the table (Intergenic, containing 12 CpG sites).
2.2 Key Feature and Highlight
- Sample Compatibility: Suitable for gDNA, cfDNA, and various level of FFPE samples.
- Multi-cancer Detection: Covers large-scale CpG sites of multiple genes for 10 major high-incidence cancers worldwide in a single detection.
- Flexible Customization: Supports tailored probe sets with various CpG site scales and/or gene mutations for single or multiple cancer types, enabling synchronous detection of methylation and mutations.
- Accurate Quantification: Achieves precise quantification of methylation levels (0 ~ 100%) in tested samples.
- Efficient Capture: Requires no sequence conversion and retains native sequence complexity, increasing effective coverage depth by several times.
- Ultra-high Sensitivity: Compatible with unique dual indexed adapters including unique molecular identifier (UMI) for ultra-sensitive detection of low to ultra-low abundance signals.
2.3 Application Expansion
• Dual-scenario Coverage for Whole-cycle
Monitoring
I. Early Screening and Risk Alerts
For target cancer types, early-screening models are constructed using cohort samples (clinical patients and healthy controls), and model risk scores are evaluated using longitudinal sample repositories to precisely assess the predictive capacity of methylation risk for transition from “precancerous lesions” to “early cancer,” thereby enabling early identification of at-risk individuals.
II. Recurrence Monitoring and Therapeutic
Efficacy Evaluation
Focusing on patients after curative surgery and/or chemotherapy/radiotherapy, periodic testing of ctDNA methylation in plasma cfDNA samples enables precise identification of MRD positivity, timely assessment of recurrence or metastasis risk, and supports treatment efficacy monitoring and personalized therapy adjustments.
• Ready-to-use or Customizable
to Flexibly Meet Personalized Needs
I. No Candidate Biomarkers
Use the μCaler FS EMS+ Panel v1.0 directly and rely on the panel’s preset key genes to rapidly initiate biomarker discovery, thereby reducing development cost and cycle time.
II. With Candidate Biomarkers
Supports personalized and flexible probe
customization (used alone or spiked in), meeting specific cancer research needs
and enabling flexible extension of application scope.
• Multi-omics Integrated Detection
Supports customized co-detection panels combining specific driver gene mutations and methylation biomarkers, markedly enhancing overall assay performance to meet personalized research and clinical exploration needs.
03 Performance
3.1 Stable Detection of
Methylation Signals
Stability of methylation detection was evaluated using tumor tissue samples from four clinically confirmed patients processed through the μCaler DNA Full Screen System workflow in combination with the μCaler FS EMS+ Panel v1.0. Each sample was tested in two technical replicates, and correlation analysis was performed on methylation levels at CpG sites included for specific cancer types in the μCaler FS EMS+ Panel v1.0. Results showed that the correlation coefficients (R²) across the four clinical samples ranged from 0.9279 to 0.9707 (Figure 1), with an average R² of 0.9458. These data indicate that the μCaler FS EMS+ Panel v1.0 maintains high stability in repeat measurements of methylation levels at the same CpG sites.
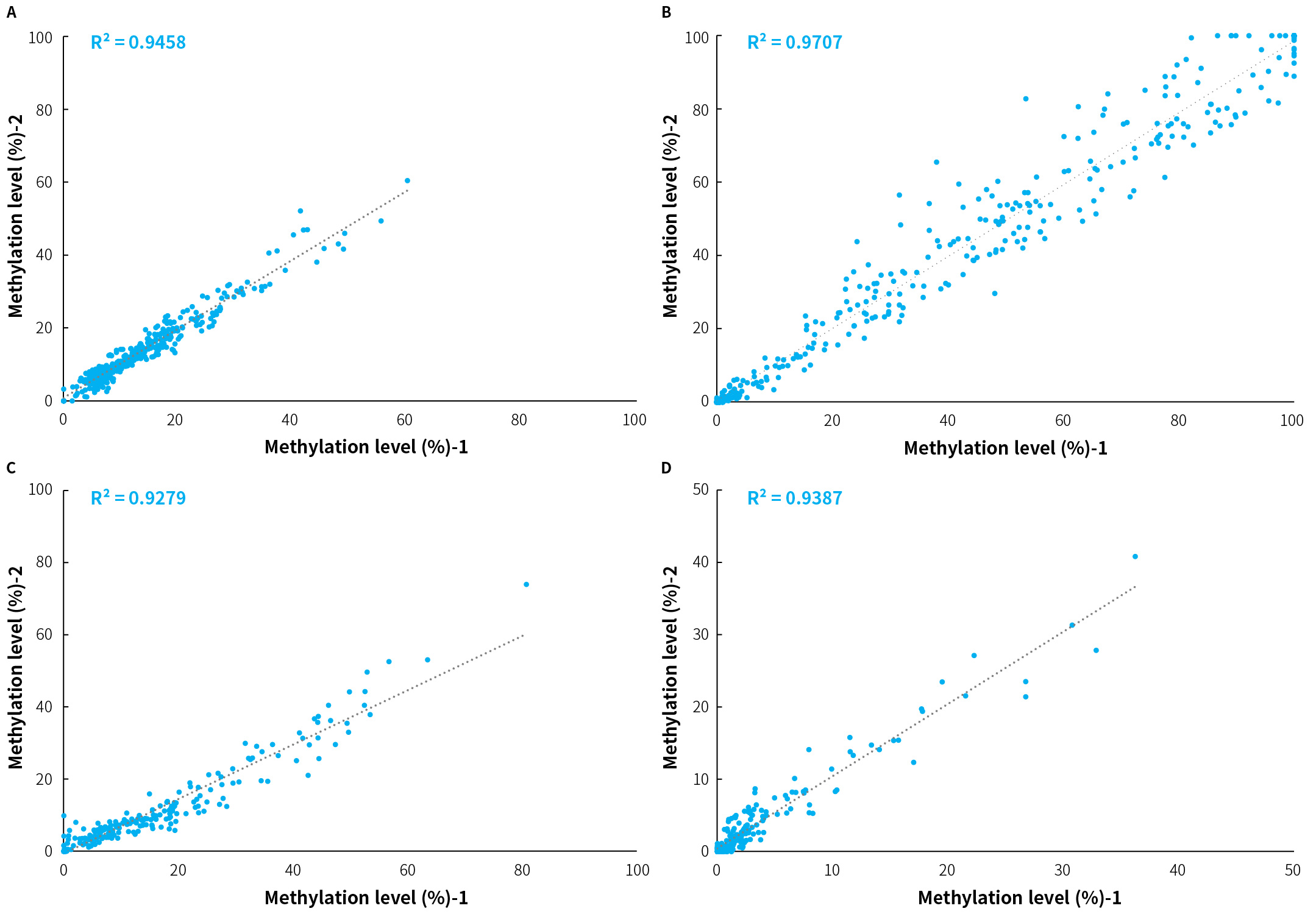
Figure 1. Correlation analysis of methylation levels detected between two repeats of the sample by μCaler FS EMS+ Panel v1.0. A-D. The correlation of methylation levels between two replicates of 4 independent samples. 4 clinical tumor tissue samples were randomly selected (A & B: colorectal cancer, C & D: bladder cancer) and processed to hybridization capture using the μCaler DNA Full Screen System and μCaler FS EMS+ Panel v1.0. The R² values for correlation ranged from 0.9279 to 0.9707.
Note: The initial input amounts were 20 ng.
3.2 Effectively Distinguishing
Tumor and Peritumoral Tissues
3.2.1 Paired Tumor and Peritumoral
Tissue (Tumor-PTT) Samples for Colorectal Cancer
The performance of colorectal cancer candidate biomarkers
in the μCaler FS EMS+ Panel v1.0 was evaluated using tumor-PTT samples from
seven clinically confirmed colorectal cancer patients and healthy controls. Results
showed that methylation levels were markedly higher in tumor tissues than in PTTs
and healthy controls (Figure 2.A); integrating methylation profiles across all
colorectal cancer candidate genes, the panel effectively distinguished tumor
tissues from PTTs and healthy controls (Figure 2.B). However, some candidate biomarkers
(e.g., SDC2) exhibited low methylation in tumor tissue of certain
patients (Figure 2.B); this observation indicates that single-gene methylation
assays may not be universally applicable, underscoring the importance of
multi-gene combined methylation analysis to improve detection sensitivity.
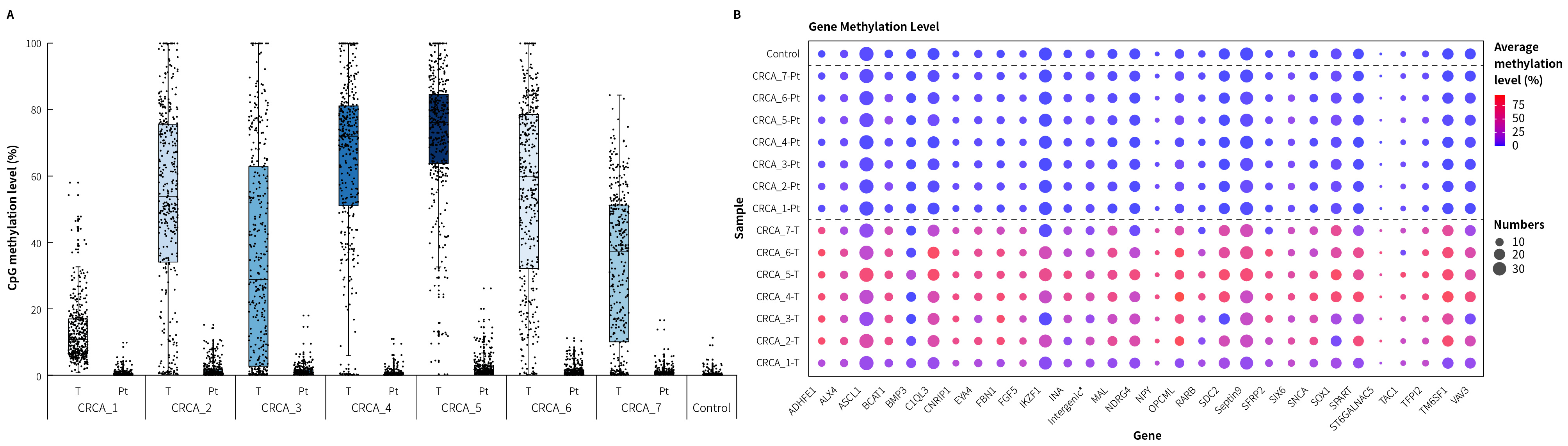
Figure 2. Methylation level detection of μCaler FS EMS+ Panel v1.0 applied to clinical colorectal cancer and peritumoral tissue samples. A. CpG methylation levels in paired colorectal cancer samples; B. CpG methylation levels of colorectal cancer candidate biomarkers in clinical samples.
Note: CRCA: Colorectal Cancer; Pt: Peritumoral tissue.
Control refers to Human Genomic DNA standard (Promega, G1471), with an input of
20 ng.
3.2.2 Tumor Tissue Samples for
Lung Cancer
The performance of lung cancer candidate biomarkers in the μCaler FS EMS+ Panel v1.0 was assessed using tumor tissue samples from five clinically confirmed lung cancer patients and healthy controls. Results showed that the five lung tumor tissues had average methylation levels of 66.09%, 21.83%, 11.10%, 22.96% and 10.87%, respectively, while the healthy controls’ average methylation level was only 0.56% (Figure 3.A); integrating methylation profiles across all lung cancer candidate genes, the panel demonstrated good discriminatory power between lung tumor tissues and healthy controls (Figure 3.B).
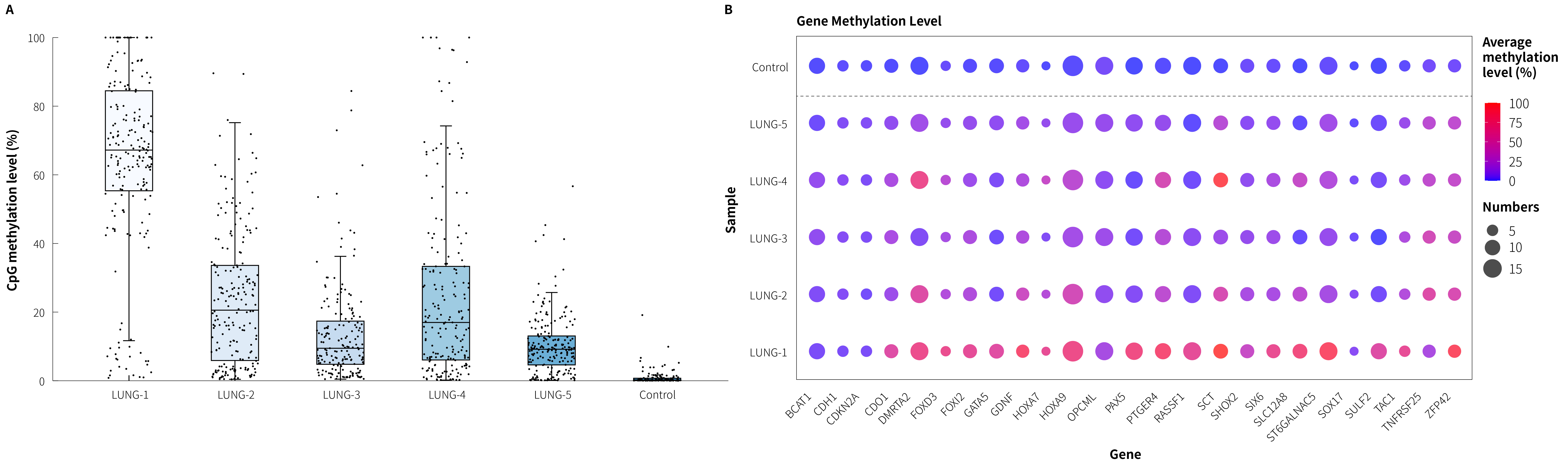
Figure 3. Methylation level detection of μCaler FS EMS+ Panel v1.0 applied to clinical lung cancer tissues. A. CpG methylation levels in lung cancer samples; B. CpG methylation levels of lung cancer candidate biomarkers in clinical samples.
Note: LUNG: Lung Cancer. The initial input amounts were 20 ng.
04 Application Examples
Scenario 1
If no large-scale DNA methylation biomarker
discovery has been performed, can one directly select preset lung cancer biomarkers
from the μCaler FS EMS+ Panel v1.0 to customize probes and use the custom panel
to evaluate its potential for lung cancer screening?
Test Results
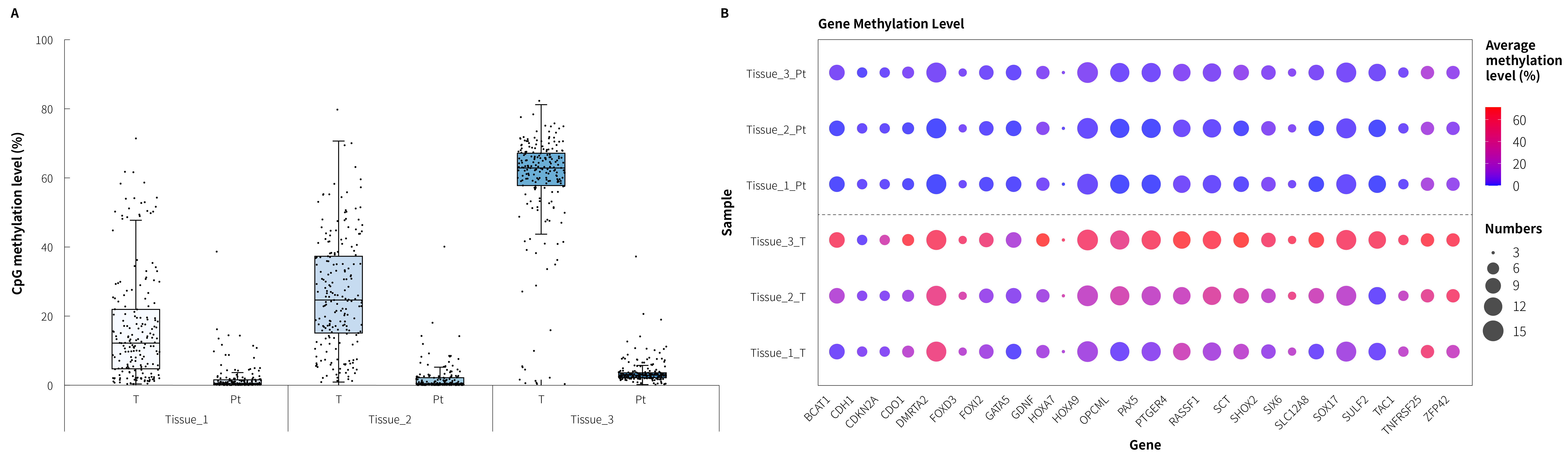
Figure 4. Methylation level detection of μCaler DNA Full Screen System applied to clinical lung cancer and peritumoral tissues. A. CpG methylation levels in paired lung cancer samples; B. CpG methylation levels of lung cancer candidate biomarkers in clinical samples. Refer to the protocol of μCaler DNA Full Screen System, hybrid capture was performed with μCaler FS customized Panel (DNA methylation: containing 24 candidate biomarkers, totaling 208 CpG sites). The sequencing mode was NovaSeq 6000, PE150. 1 M reads pair were randomly selected for data analysis.
Note: T: cancer tissue; Pt: peritumoral tissue. The initial input amounts were 50 ng.
The performance of colorectal cancer candidate biomarkers in the μCaler FS customized panel was evaluated using tumor-PTT samples from three clinically confirmed lung cancer patients. Results showed that methylation levels were markedly higher in tumor tissues than in PTTs and healthy controls (Figure 4.A); integrating methylation profiles across all lung cancer candidate genes, the customized panel effectively distinguished tumor tissues from PTTs (Figure 4.B). This observation indicates that that the preset lung cancer biomarkers in the μCaler FS EMS+ Panel v1.0 have potential utility for lung cancer screening and can support discovery of biomarkers and construction of screening models.
Application Summary
When no in-house candidate biomarkers exist,
investigators can freely select preset biomarkers for single or multiple
cancers from the μCaler FS EMS+ Panel v1.0 to rapidly customize a panel,
perform methylation testing on clinical cohort samples, and construct an
early-screening model for the target cancer. Subsequently, use the longitudinal
sample repository to score the model and evaluate the ability of methylation
risk score to predict the window of transition from “precancerous lesions” to “early
cancer”, and to define risk stratification and alert thresholds for early
warning of clinical samples.
Aberrant DNA methylation is a key epigenetic
modification driving cancer initiation and progression, and it often occurs in
early stages of tumorigenesis. Therefore, DNA methylation can serve as a
biomarker for early cancer detection and prognostic assessment, and it is
valuable for screening and risk evaluation, early diagnosis, staging,
prognostication and treatment monitoring.
Scenario 2
If lung cancer-relevant mutation loci have already
been identified, can they be combined with lung cancer biomarkers selected from
the μCaler FS EMS+ Panel v1.0 to synthesize a single-tube probe set, and can
this custom panel be used to evaluate the potential of combined DNA methylation
and mutation biomarkers for MRD recurrence monitoring in lung cancer?
Test results
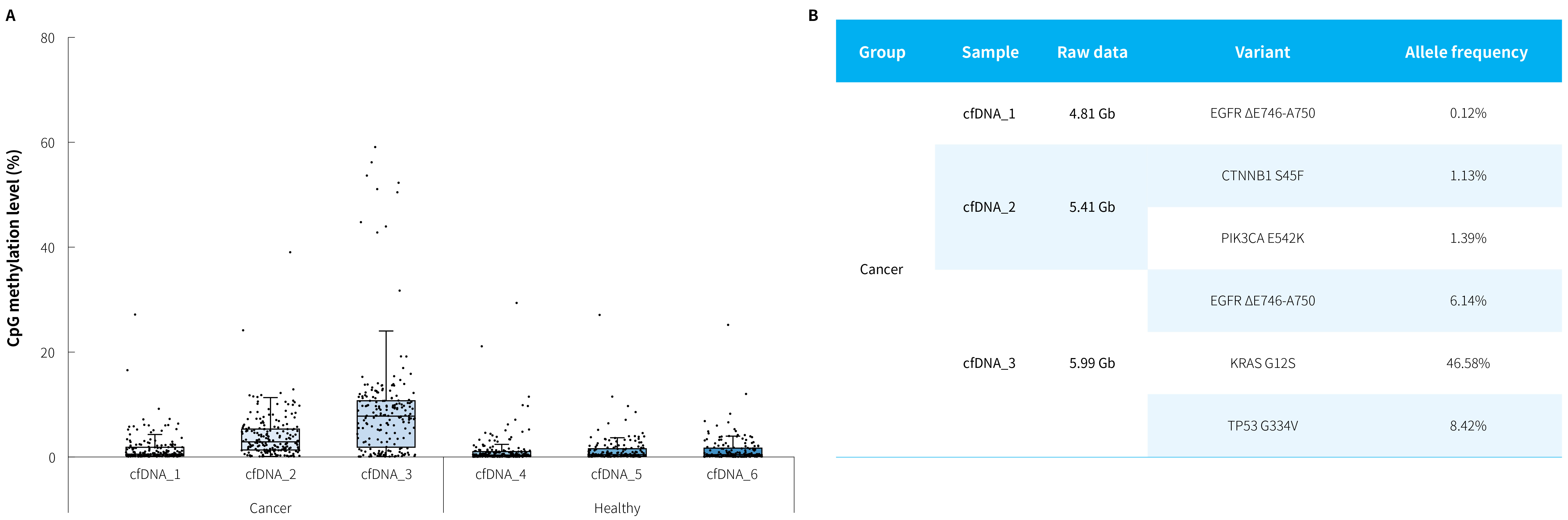
Figure 5. Capture performance of μCaler DNA Full Screen System applied to MRD recurrence monitoring in clinical lung cancer patients. A. CpG methylation levels of lung cancer candidate biomarkers in cfDNA samples from clinical post-operative recurrence patients and healthy individuals; B. Mutation frequencies of lung cancer candidate biomarkers in cfDNA samples from clinical post-operative recurrence patients. Refer to the protocol of μCaler DNA Full Screen System, hybrid capture was performed with μCaler FS customized Panel (DNA methylation: containing 24 candidate biomarkers, totaling 208 CpG sites; DNA mutation: containing 5 candidate biomarkers). The sequencing mode was NovaSeq 6000, PE150. The variant analysis was performed using fgbio in combination with Vardict [pre-libraries containing UMIs, with analytical filtering criteria set to duplex consensus sequences (DCS211)].
Note: Cancer: plasma cfDNA from lung cancer recurrence patients; Healthy: plasma
cfDNA from healthy population. The initial input amounts were 20 ng.
Using plasma cfDNA from three postoperative lung
cancer recurrence patients and three healthy controls, the μCaler FS customized
panel’s performance for MRD monitoring was evaluated. Results indicated average
methylation levels of 1.4%, 3.8% and 8.7% in the recurrence patients,
respectively, showing a gradient increase with recurrence progression; compared
with healthy controls, MRD positive status could be discriminated (Figure 5.A).
Concurrently, all three recurrence patients’ cfDNA samples contained
corresponding MRD-specific mutations (Figure 5.B), providing strong mutual
validation with the methylation biomarker results.
Application Summary
Based on candidate mutation biomarkers, methylation
biomarkers can be freely chosen from the μCaler FS EMS+ Panel v1.0 to
rapidly synthesize a single-tube probe set, which-together with the μCaler DNA
Full Screen System—can be used to establish a stable “no-disease” baseline
methylation and/or mutation profiles from the patients’ postoperative samples.
Define alert thresholds based on baseline distributions and use the panel to
monitor cfDNA methylation level in serial follow-up samples to determine MRD
positivity or trigger recurrence alerts. Continuous monitoring during
subsequent treatments allows assessment of therapeutic response and guides
personalized therapy.
Compared with mutation-only monitoring, methylation biomarkers do not require knowledge of the primary tumor’s mutation profile and are not limited by mutation frequency or the number of mutation loci, thereby substantially improving MRD monitoring sensitivity and specificity. Integrating methylation and mutation features enables multidimensional coverage from genomic alterations to epigenetic regulation, considerably enhancing MRD detection accuracy and clinical decision-making value.
05 Future Prospects
In precision oncology, DNA methylation biomarkers
play an increasingly pivotal role in early detection, treatment selection and
prognostic assessment. Efficiently screening and validating methylation biomarkers
to build early-screening models has become a current research focus. To meet
this need, Nanodigmbio has launched a ready-to-use methylation detection panel
covering 10 major high-incidence cancers: the μCaler FS EMS+ Panel v1.0. Based
on methylation-sensitive restriction endonuclease (MSRE) principles and
combined with μCaler patented probe design, and deployed on the μCaler DNA Full
Screen System, the panel is applicable to cancer screening and MRD
surveillance. Users may flexibly use the panel’s preset candidate biomarkers or
customize probes to meet single-cancer or multi-cancer research needs,
shortening development timelines and accelerating clinical translation.
It should be emphasized that, due to the nature of
clinical research and translational application, cancer screening and MRD
surveillance are inherently evolving processes without a fixed end point.
Moving forward, Nanodigmbio will continue to follow the latest advances in
oncology research and leverage its diversified technology platforms and
comprehensive service capabilities to provide end-to-end customized solutions. Nanodigmbio
aims to fully support technology and product development across the entire
workflow of cancer screening, diagnosis and treatment monitoring, collaborating
to advance cancer control and management and contribute to the development of
precision oncology.
Reference
[1] Palanca-Ballester C,
Rodriguez-Casanova A, Torres S, et al. Cancer epigenetic biomarkers in liquid
biopsy for high incidence malignancies[J]. Cancers, 2021, 13(12): 3016.
[2] Ibrahim J, Peeters M, Van Camp G, et
al. Methylation biomarkers for early cancer detection and diagnosis: Current
and future perspectives[J]. European Journal of Cancer, 2023, 178: 91-113.
[3] Jin S, Zhu D, Shao F, et al.
Efficient detection and post-surgical monitoring of colon cancer with a
multi-marker DNA methylation liquid biopsy[J]. Proceedings of the National
Academy of Sciences, 2021, 118(5): e2017421118.
[4] Li F, Zheng Z, Chen W, et al.
Regulation of cisplatin resistance in bladder cancer by epigenetic
mechanisms[J]. Drug Resistance Updates, 2023, 68: 100938.
[5] Ohnmacht A J, Rajamani A, Avar G, et
al. The pharmacoepigenomic landscape of cancer cell lines reveals the
epigenetic component of drug sensitivity[J]. Communications Biology, 2023,
6(1): 825.
[6] Parikh A R, Van Seventer E E, Siravegna G, et al. Minimal residual disease detection using a plasma-only circulating tumor DNA assay in patients with colorectal cancer[J]. Clinical Cancer Research, 2021, 27(20): 5586-5594.
Solutions
- Methyl Library Preparation Total Solution
- Sequencing single library on different platform--Universal Stubby Adapter (UDI)
- HRD score Analysis
- Unique Dual Index for MGI platforms
- RNA-Cap Sequencing of Human Respiratory Viruses Including SARS-CoV-2
- Total Solution for RNA-Cap Sequencing
- Total Solution for MGI Platforms
- Whole Exome Sequencing
- Low-frequency Mutation Analysis
Events
-
Exhibition Preview | Nanodigmbio invites you to join us at Boston 2025 Annual Meeting of the American Society of Human Genetics (ASHG)

-
Exhibition Preview | Nanodigmbio Invites You to Join Us at WHX & WHX Labs Kuala Lumpur 2025, Malaysia International Trade and Exhibition Centre in Kuala Lumpur

-
Exhibition Preview | Nanodigmbio Invites You to Join Us at Hospitalar 2025, Brazil International Medical Device Exhibition in São Paulo

-
Exhibition Preview | Nanodigmbio invites you to join us at Denver 2024 Annual Meeting of the American Society of Human Genetics (ASHG)

-
Exhibition Preview | Nanodigmbio invites you to join us at Sapporo 2024 Annual Meeting of the Japan Society of Human Genetics (JSHG)

-
Exhibition Preview | Nanodigmbio invites you to join us at Association for Diagnostics & Laboratory Medicine (ADLM)


