μCaler RIA Solution: Unlocking a New Paradigm of Precision Diagnosis and Treatment for Respiratory Tract Infections
In our previous article, we focused on the core challenges in the diagnosis and treatment of respiratory tract infections, highlighting the inevitable transition from empirical medication to precision diagnosis and treatment, and systematically introduced the overall design philosophy of μCaler RIA Solution in multiplex pathogen detection, antimicrobial resistance(AMR) profiling, and result interpretation.
On this basis, it becomes clear that in real-world clinical practice, the critical question is whether improved detection capabilities can truly translated into clinical value? In other words, an effective diagnostic solution must not only “identify pathogens,” but also assist clinical judgment, guide therapeutic decisions, and provide actionable decision support in complex infection scenarios. Therefore, the evaluation of a diagnostic solution should consider not only its technical performance, but also its real-world clinical utility and its ability to support clinical decision-making. How, then, does μCaler RIA Solution perform in respiratory tract infection diagnostics? This article further elaborates on its detection performance and AMR profiling capabilities.
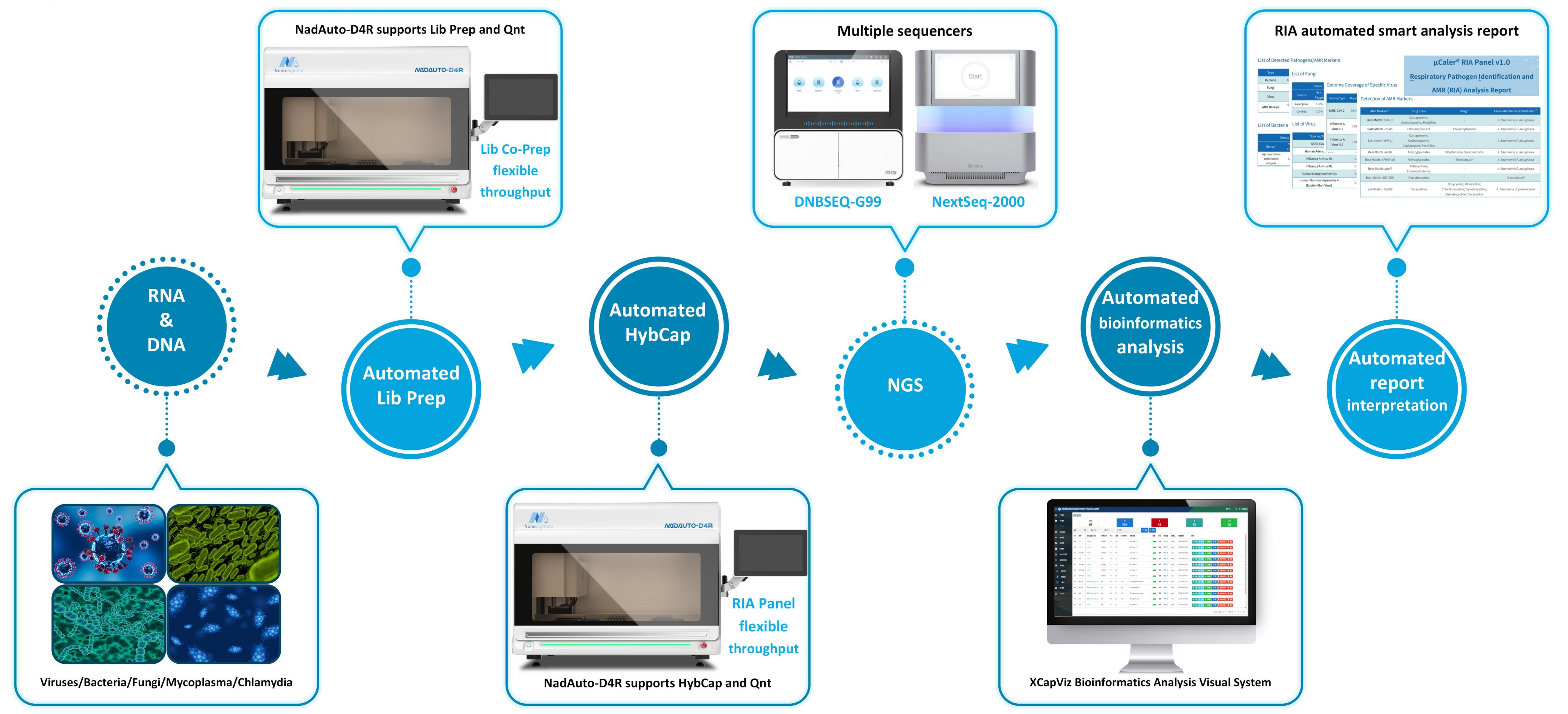
Figure 1. Overview of μCaler RIA Solution.
01 Stable Detection of Multiple Pathogens in Reference Standard
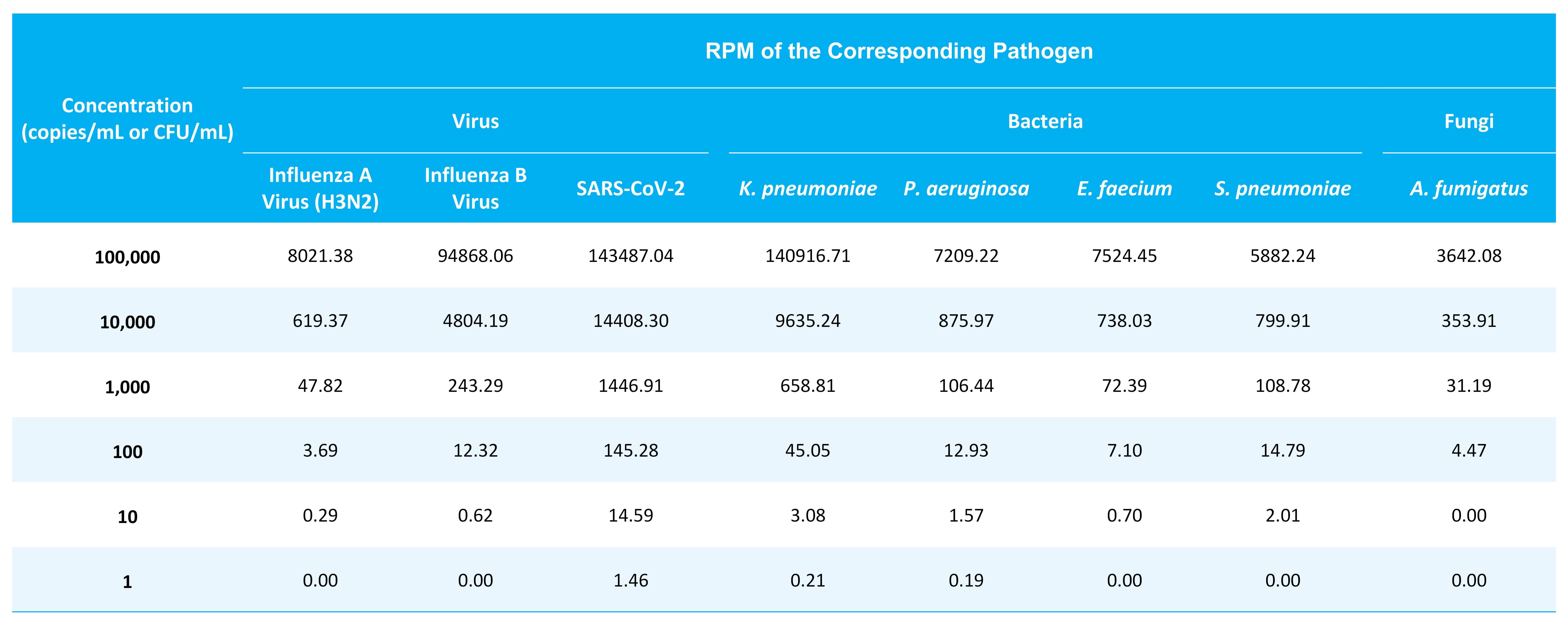
To systematically evaluate the detection performance of μCaler RIA Solution, microbial reference standards were serially diluted to assess detection sensitivity across different concentrations for representative pathogens, including Influenza A virus (H3N2), SARS-CoV-2, K. pneumoniae, S. pneumoniae, E. faecium, A. fumigatus. Specifically, a total of 0.4 mL of reference standards at different concentration gradients were used for extraction of mixed RNA & DNA. Pre-libraries were prepared using the NadPrep EZ RNA & DNA Library Co-Preparation Module coupled with the NadPrep Universal Stubby Adapter (UDI) Module. Hybrid capture was performed using the μCaler HyperMix Hybrid Capture Kit (for Illumina) and the μCaler RIA Panel v1.0. RPM (reads per million mapped reads)-based analysis demonstrated that, among the 6 pathogens across 3 categories, all pathogens except A. fumigatus could be consistently detected at concentrations as low as 10-100 copies/mL or 10-100 CFU/mL. Notably, owing to the whole-genome coverage design for SARS-CoV-2 in μCaler RIA Panel v1.0, the detection sensitivity for this virus reached as low as 1-10 genome copies (Table 1).
Table 1. Limits of Detection (LOD) of μCaler RIA Solution for Multiplex Pathogens in Reference Materials.
In addition, the reproducibility of the μCaler RIA Solution was evaluated using reference standards across both intra-run (within-batch) and inter-run (between-batch) conditions. Based on RPM analysis, the solution achieved stable detection across different batches for the multi-type pathogens (including viruses, bacteria, and fungi) included in the reference standards. In addition, results showed good consistency within the same batch (Figure 2). These findings indicate that the μCaler RIA Solution delivers excellent reproducibility and reliability for multiplex pathogen detection, supporting high-throughput clinical testing with robust performance.
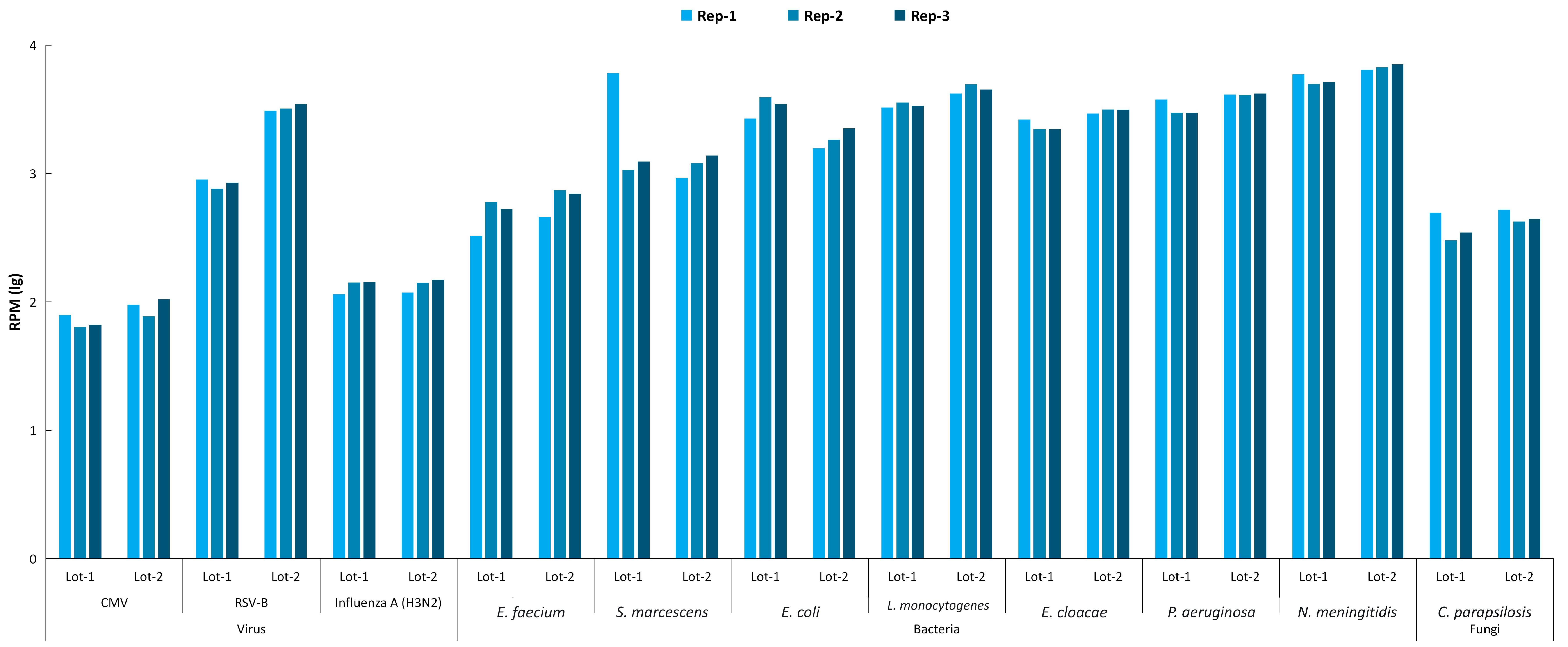
Figure 2. Intra- and inter-batch stability evaluation of μCaler RIA Solution for multiplex pathogen detection in reference standards.
02 Enhanced Detection Rates for Clinically Critical Pathogens
To further assess its clinical performance, we conducted an integrated analysis of samples from 78 clinical patients (including multiple specimen types such as sputum and BALF) to compare the μCaler RIA Solution with conventional routine clinical testing across six key pathogens. The findings show that the overall detection rates of both μCaler RIA Solution and mNGS were significantly higher than culture (Figure 3). The advantage was especially pronounced for P. jirovecii and Nocardia, largely because these pathogens are not readily culturable in vitro, limiting culture-based detection. At the individual pathogen level, the μCaler RIA Solution detected more cases than mNGS for the MTB complex (MTBC), P. jirovecii, and A. fumigatus. In contrast, mNGS detected more cases than μCaler RIA Solution for NTM and Nocardia. These results suggest that the two technologies offer complementary strengths across different pathogen types.
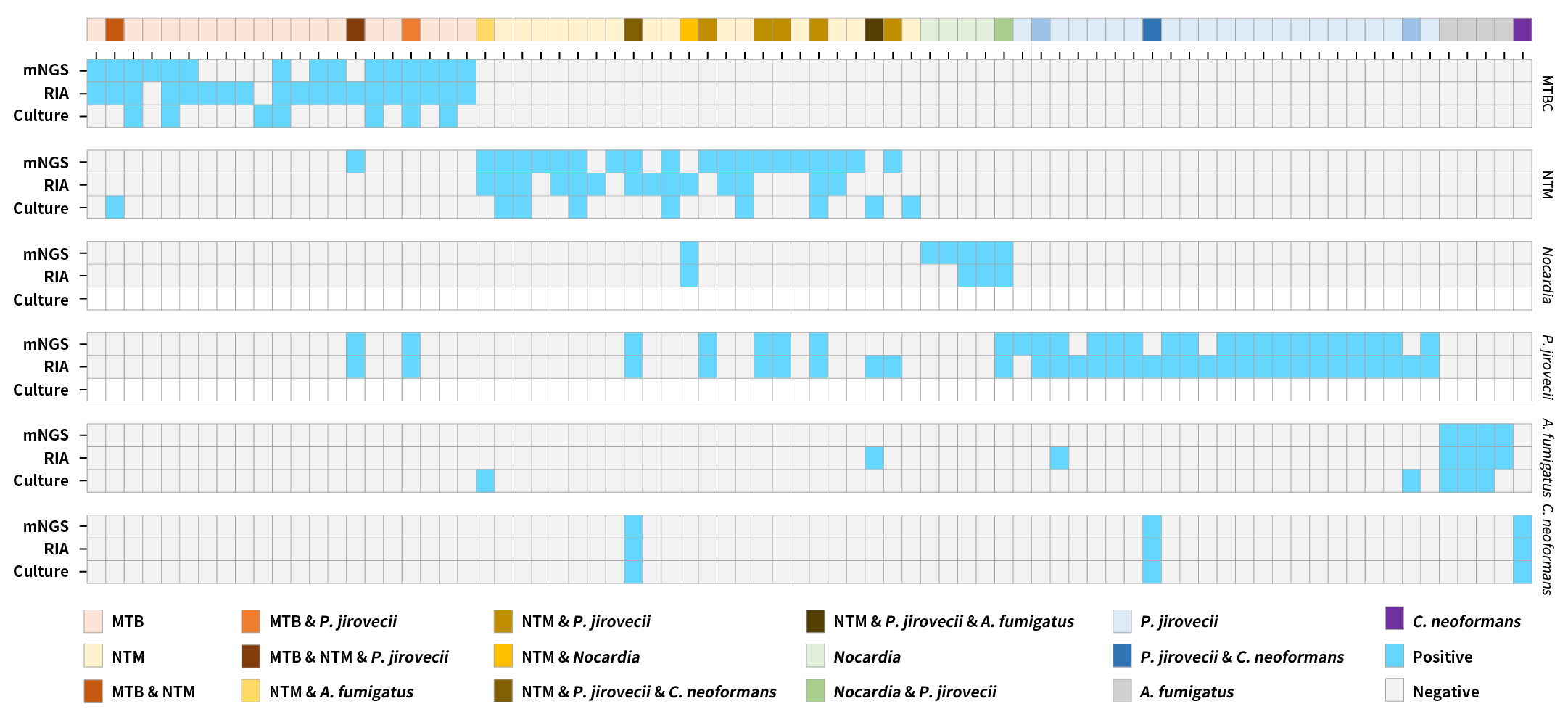
Figure 3. Heatmap comparison of detection concordance among the μCaler RIA Solution, mNGS, and culture for key clinical pathogens.
Note:Light blue and light gray blocks indicate positive and negative results, respectively; white block indicates the method is not applicable, and no valid result could be obtained. Clinically important pathogens included in the analysis were: MTBC (M. tuberculosis complex), NTM (nontuberculous mycobacteria), Nocardia spp., P. jirovecii, A. fumigatus, and C. neoformans.
03 Precise Identification of Antimicrobial Resistance Genes (ARGs) and Associated Pathogens
To systematically evaluate AMR profiling capability, detection results from the μCaler RIA Solution were compared with conventional microbiological methods [including culture-based identification and phenotypic antimicrobial susceptibility testing (AST)] in three clinical cases. The results showed high concordance in pathogen identification, detecting K. pneumoniae, P. aeruginosa, and A. baumannii respectively. Furthermore, the ARGs associated antimicrobial classes analyzed by the μCaler RIA Solution were highly consistent with the phenotypes of AST (Table 2). These findings indicate that μCaler RIA Solution can accurately associate resistance genes with their corresponding pathogens and infer potential antimicrobial resistance, thereby providing critical molecular evidence to guide clinical treatment decisions and enhance clinical value of respiratory tract infection diagnosis and treatment.
Table 2. Concordance analysis between the μCaler RIA Solution and conventional methods for pathogens detection and distribution of ARGs across diverse clinical specimens.
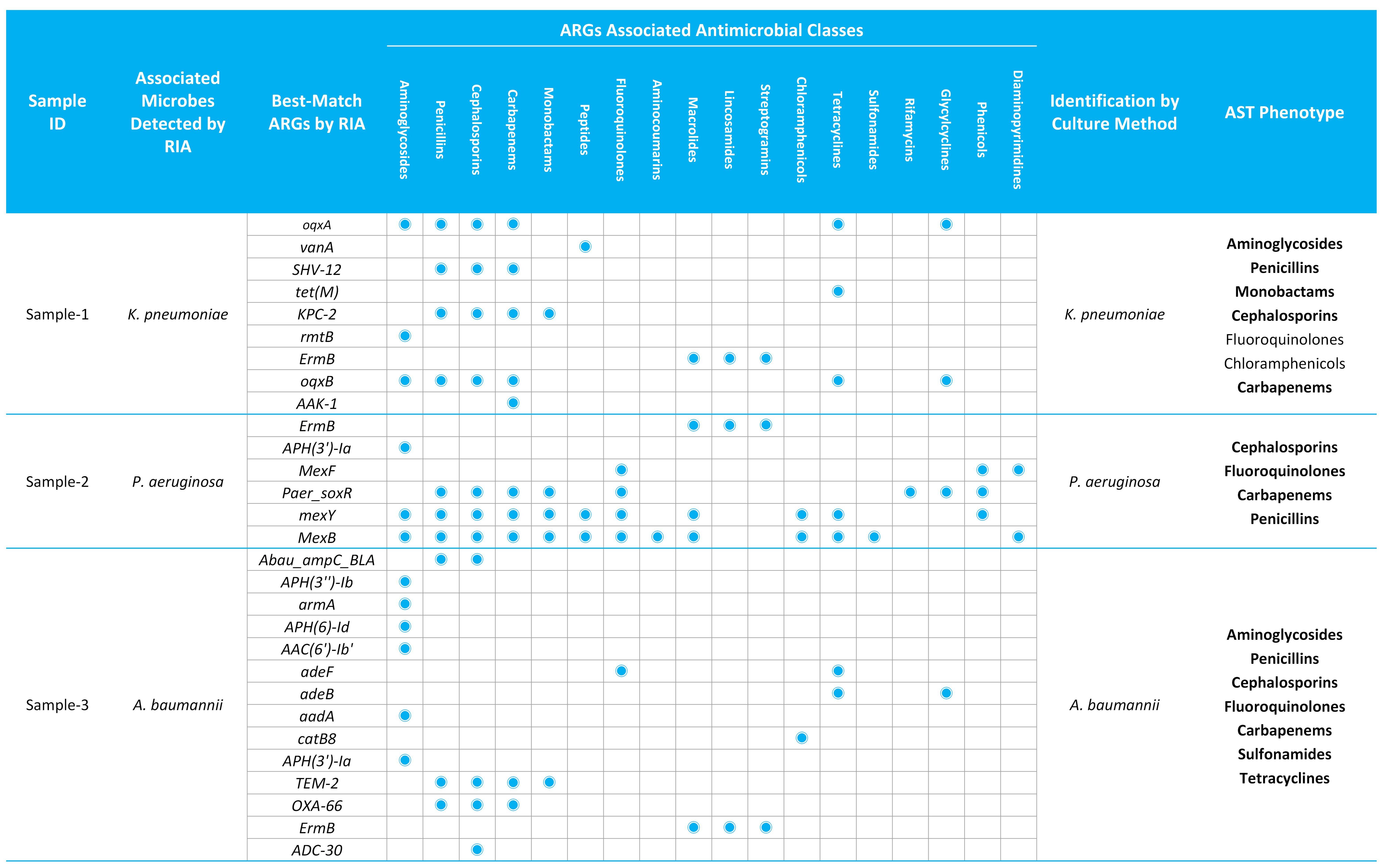
Note: ARGs: Antibiotic Resistance Genes.
04 Summary and Outlook
Addressing the core pain points of "difficulty in detection, interpretation, and decision-making" in respiratory tract infection diagnosis, μCaler RIA Solution delivers an integrated workflow covering multiplex pathogen detection, AMR profiling, and intelligent result interpretation. Evaluations using both reference standards and clinical samples demonstrate its strong performance in sensitivity, reproducibility, and robustness, enabling reliable identification of low-abundance pathogens in infection scenarios. Moreover, through precise detection of ARGs and their linkage to source pathogens, the solution effectively translates molecular findings into actionable clinical guidance. Combined with a hierarchical interpretation framework and multi-dimensional quality control system, it further improves the interpretability and clinical reference of diagnostic results.
Going forward, as the spectrum of respiratory pathogens continues to evolve and AMR becomes increasingly severe, the demand for diagnostic solutions that are "rapid, comprehensive, precise, and interpretable" will continue to grow. The μCaler RIA Solution is well-positioned to play a pivotal role in broader range of clinical applications. With ongoing optimization of algorithms and database systems, its capabilities in result interpretation and clinical decision support are expected to further improve, facilitating the transition of respiratory tract infection management from empirically driven approaches to data-driven precision medicine.
Solutions
- Methyl Library Preparation Total Solution
- Sequencing single library on different platform--Universal Stubby Adapter (UDI)
- HRD score Analysis
- Unique Dual Index for MGI platforms
- RNA-Cap Sequencing of Human Respiratory Viruses Including SARS-CoV-2
- Total Solution for RNA-Cap Sequencing
- Total Solution for MGI Platforms
- Whole Exome Sequencing
- Low-frequency Mutation Analysis
Events
-
Exhibition Preview | Nanodigmbio invites you to join us at Boston 2025 Annual Meeting of the American Society of Human Genetics (ASHG)

-
Exhibition Preview | Nanodigmbio Invites You to Join Us at WHX & WHX Labs Kuala Lumpur 2025, Malaysia International Trade and Exhibition Centre in Kuala Lumpur

-
Exhibition Preview | Nanodigmbio Invites You to Join Us at Hospitalar 2025, Brazil International Medical Device Exhibition in São Paulo

-
Exhibition Preview | Nanodigmbio invites you to join us at Denver 2024 Annual Meeting of the American Society of Human Genetics (ASHG)

-
Exhibition Preview | Nanodigmbio invites you to join us at Sapporo 2024 Annual Meeting of the Japan Society of Human Genetics (JSHG)

-
Exhibition Preview | Nanodigmbio invites you to join us at Association for Diagnostics & Laboratory Medicine (ADLM)


